Nonlinear regression plays an important role in my research and teaching activities. While I often use the ‘drm()’ function in the ‘drc’ package for my research work, I tend to prefer the ‘nls()’ function for teaching purposes, mainly because, in my opinion, the transition from linear models to nonlinear models is smoother, for beginners. One problem with ‘nls()’ is that, in contrast to ‘drm()’, it is not specifically tailored to the needs of biologists or students in biology. Therefore, now and then, I have to build some helper functions, to perform some specific tasks; I usually share these functions within the ‘aomisc’ package, that is available on github (see this link).
In this post, I would like to describe one of these helper functions, i.e. ‘gnlht()’, which is aimed at helping students (and practitioners; why not?) with one of their tasks, i.e. making some simple manipulations of model parameters, to obtain relevant biological information. Let’s see a typical example.
Motivating example
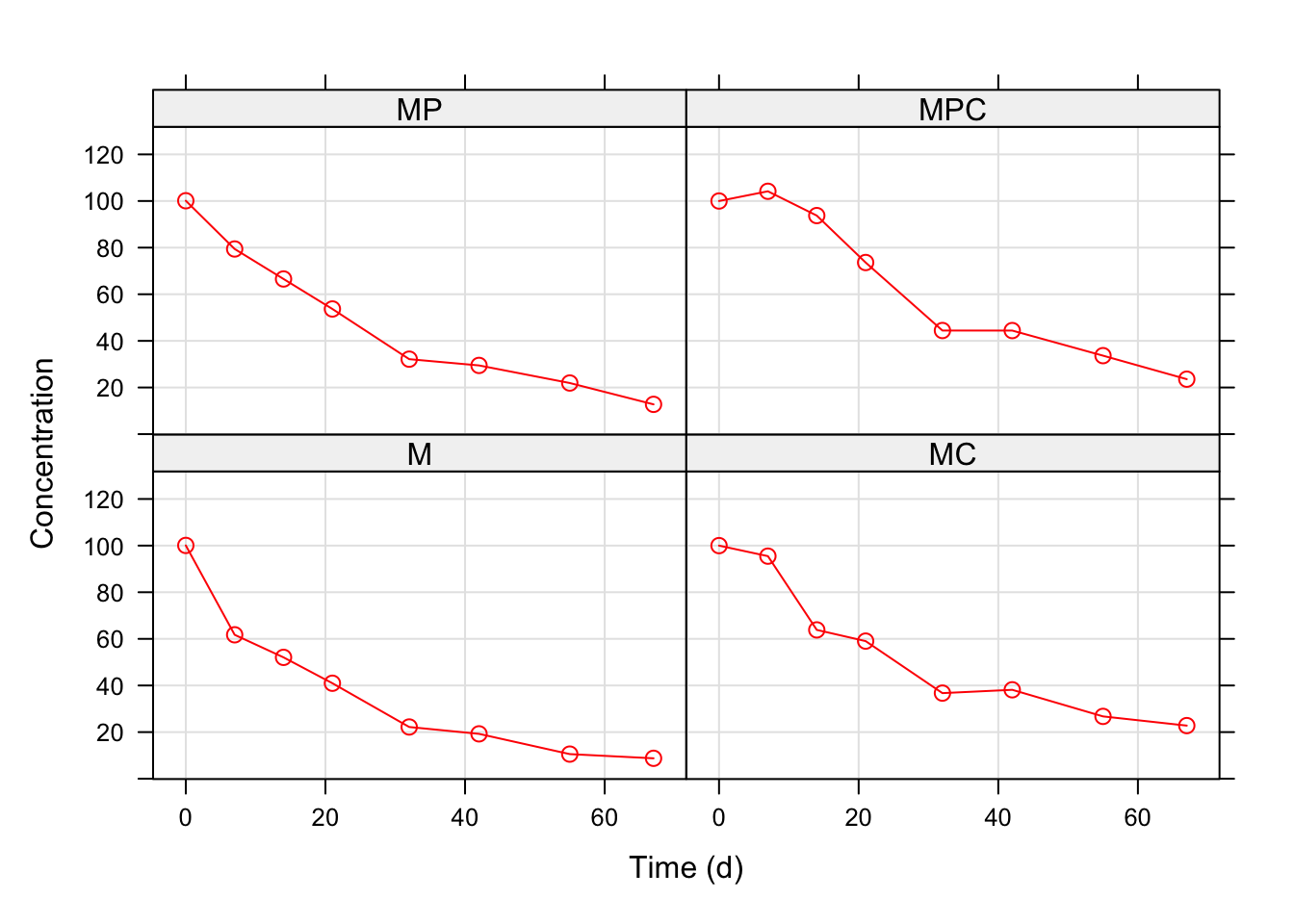
This is a real-life example, taken from a research published by Vischetti et al. in 1996. That research considered three herbicides for weed control in sugar beet, i.e. metamitron (M), phenmedipham (P) and cloridazon (C). Four soil samples were contaminated, respectively with: (i) M alone, (ii) M + P (iii) M + C and (iv) M + P + C. The aim was to assess whether the degradation speed of metamitron in soil depended on the presence of co-applied herbicides. To reach this aim, the soil samples were incubated at 20°C and sub-samples were taken in different times after the beginning of the experiment. The concentration of metamitron in those sub-samples was measured by HPLC analyses, performed in triplicate. The resulting dataset is available within the ‘aomisc’ package; we can load it and use the ‘lattice’ package to visualise the observed means over time.
# library(devtools)
# install_github("OnofriAndreaPG/aomisc")
library(aomisc)
library(lattice)
data(metamitron)
xyplot(Conc ~ Time|Herbicide, data = metamitron,
xlab = "Time (d)", ylab = "Concentration",
scales = list(alternating = F),
panel = function(x, y, ...) {
panel.grid(h = -1, v = -1)
fmy <- tapply(y, list(factor(x)), mean)
fmx <- tapply(x, list(factor(x)), mean)
panel.xyplot(fmx, fmy, col="red", type="b", cex = 1)
})
Considering this exemplary dataset, let’s see how we can answer the following research questions.
- What is the degradation rate for metamitron, in the four combinations?
- Is there a significant difference between the degradation rate of metamitron alone and with co-applied herbicides?
- What is the half-life for metamitron, in the four combinations?
- What are the times to reach 70 and 30% of the initial concentration, for metamitron in the four combinations?
- Is there a significant difference between the half-life of metamitron alone and with co-applied herbicides?
Fitting a degradation model
The figure above shows a visible difference in the degradation pattern of metamitron, which could be attributed to the presence of co-applied herbicides. The degradation kinetics can be described by the following (first-order) model:
\[ C(t, h) = A_h \, \exp \left(-k_h \, t \right) \]
where \(C(t, h)\) is the concentration of metamitron at time \(t\) in each of the four combinations \(h\), \(A_h\) and \(k_h\) are, respectively, the initial concentration and degradation rate for metamitron in each combination.
The model is nonlinear and, therefore, we can use the ‘nls()’ function for nonlinear least squares regression. The code is given below: please, note that the two parameters are followed by the name of the factor variable in square brackets (i.e.: A[Herbicide] and k[Herbicide]). This is necessary to fit a different parameter value for each level of the ‘Herbicide’ factor.
#Fit nls grouped model
modNlin <- nls(Conc ~ A[Herbicide] * exp(-k[Herbicide] * Time),
start=list(A=rep(100, 4), k=rep(0.06, 4)),
data=metamitron)
summary(modNlin)
##
## Formula: Conc ~ A[Herbicide] * exp(-k[Herbicide] * Time)
##
## Parameters:
## Estimate Std. Error t value Pr(>|t|)
## A1 9.483e+01 4.796e+00 19.77 <2e-16 ***
## A2 1.021e+02 4.316e+00 23.65 <2e-16 ***
## A3 9.959e+01 4.463e+00 22.31 <2e-16 ***
## A4 1.116e+02 4.184e+00 26.68 <2e-16 ***
## k1 4.260e-02 4.128e-03 10.32 <2e-16 ***
## k2 2.574e-02 2.285e-03 11.26 <2e-16 ***
## k3 3.034e-02 2.733e-03 11.10 <2e-16 ***
## k4 2.186e-02 1.822e-03 12.00 <2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 9.701 on 88 degrees of freedom
##
## Number of iterations to convergence: 5
## Achieved convergence tolerance: 7.149e-06For the sake of simplicity, I will neglige an accurate model check, although I need to point out that this is highly wrong. I’ll come back to this issue in another post.
Working with model parameters
Considering the research questions, it is clear that the above output answers the first one, as it gives the four degradation rates, \(k1\), \(k2\), \(k3\) and \(k4\). All the other questions can be translated into sets of linear/nonlinear functions (combinations) of model parameters. If we use the naming of parameter estimates in the nonlinear regression object, for the second question we can write the following functions: \(k1 - k2\), \(k1 - k3\) and \(k1 - k4\). The third question requires some slightly more complex math: if we invert the equation above for one herbicide, we get to the following inverse:
\[ t = \frac{- log \left[\frac{C(t)}{A} \right] }{k} \]
I do not think this is complex enough to scare the biologists, is it? The half-life is the time required for C(t) to drop to half of the initial value, so that \(C(t)/A\) is equal to \(0.5\). Thus:
\[ t_{1/2} = \frac{- \log \left[0.5 \right] }{k} \]
Analogously, we can answer the question 4, by replacing \(0.5\) respectively with \(0.7\) and \(0.3\). The difference between the half-lives of metamitron alone and combined with the second herbicide can be calculated by:
\[ \frac{- \log \left[0.5 \right] }{k_1} - \frac{- \log \left[0.5 \right] }{k_2} = \frac{k_2 - k_1}{k_1 \, k_2} \, \log(0.5)\]
The other differences are obtained analogously.
Inferences and hypotheses testing
All parameter estimates are characterised by some uncertainty, which is summarised by way of the standard errors (see the code output above). Clearly, such an uncertainty propagates to their combinations. As for the first question, the combinations are linear, as only subtraction is involved. Therefore, the standard error for the difference can be easily calculated by the usual law of propagation of errors, which I have dealt with in this post.
In R, linear combinations of model parameters can be built and tested by using the ‘glht()’ function in the ‘multcomp’ package. However, I wanted to find a general solution, that could handle both linear and nonlinear combinations of model parameters. Such a solution should be based on the ‘delta method’, which I have dealt with in this post. Unfortunately, the function ‘deltaMethod()’ in the ‘car’ package is not flexible enough to the aims of my students and mine.
Therefore, I wrote a wrapper for the ‘deltaMethod()’ function, which I named ‘gnlht()’, as it might play for nonlinear combinations the same role as ‘glht()’ for linear combinations. To use this function, apart from loading the ‘aomisc’ package, we need to prepare a list of formulas. Care needs to be taken to make sure that the element in the formulas correspond to the names of the estimated parameters in the model object, as returned by the ‘coef()’ method. In the box below, I show how we can calculate the differences between the degradation rates.
funList <- list(~k1 - k2, ~k1 - k3, ~k1 - k4)
gnlht(modNlin, funList)
## Form Estimate SE t-value p-value
## 1 k1 - k2 0.01686533 0.004718465 3.574325 5.727310e-04
## 2 k1 - k3 0.01226241 0.004951372 2.476568 1.517801e-02
## 3 k1 - k4 0.02074109 0.004512710 4.596150 1.430392e-05The very same code can be used for nonlinear combinations of model parameters. In order to calculate the half-lives, we can use the following code:
funList <- list(~ -log(0.5)/k1, ~ -log(0.5)/k2,
~ -log(0.5)/k3, ~ -log(0.5)/k4)
gnlht(modNlin, funList)
## Form Estimate SE t-value p-value
## 1 -log(0.5)/k1 16.27089 1.576827 10.31876 7.987828e-17
## 2 -log(0.5)/k2 26.93390 2.391121 11.26413 9.552912e-19
## 3 -log(0.5)/k3 22.84747 2.058588 11.09861 2.064093e-18
## 4 -log(0.5)/k4 31.70942 2.643330 11.99601 3.257068e-20Instead of writing ‘0.5’, we can introduce a new model term, e.g. ‘prop’, as a ‘constant’, in the sense that it is not an estimated parameter. We can pass a value for this constant in a data frame, by using the ‘const’ argument:
funList <- list(~ -log(prop)/k1, ~ -log(prop)/k2,
~ -log(prop)/k3, ~ -log(prop)/k4)
gnlht(modNlin, funList, const = data.frame(prop = 0.5))
## Form prop Estimate SE t-value p-value
## 1 -log(prop)/k1 0.5 16.27089 1.576827 10.31876 7.987828e-17
## 2 -log(prop)/k2 0.5 26.93390 2.391121 11.26413 9.552912e-19
## 3 -log(prop)/k3 0.5 22.84747 2.058588 11.09861 2.064093e-18
## 4 -log(prop)/k4 0.5 31.70942 2.643330 11.99601 3.257068e-20This is very flexible, because it lets us to calculate, altogether, the half-life and the times required for the concentration to drop to 70 and 30% of the initial value:
funList <- list(~ -log(prop)/k1, ~ -log(prop)/k2,
~ -log(prop)/k3, ~ -log(prop)/k4)
gnlht(modNlin, funList, const = data.frame(prop = c(0.7, 0.5, 0.3)))
## Form prop Estimate SE t-value p-value
## 1 -log(prop)/k1 0.7 8.372564 0.8113927 10.31876 7.987828e-17
## 2 -log(prop)/k2 0.7 13.859465 1.2304069 11.26413 9.552912e-19
## 3 -log(prop)/k3 0.7 11.756694 1.0592942 11.09861 2.064093e-18
## 4 -log(prop)/k4 0.7 16.316815 1.3601865 11.99601 3.257068e-20
## 5 -log(prop)/k1 0.5 16.270892 1.5768267 10.31876 7.987828e-17
## 6 -log(prop)/k2 0.5 26.933905 2.3911214 11.26413 9.552912e-19
## 7 -log(prop)/k3 0.5 22.847468 2.0585881 11.09861 2.064093e-18
## 8 -log(prop)/k4 0.5 31.709416 2.6433295 11.99601 3.257068e-20
## 9 -log(prop)/k1 0.3 28.261979 2.7388938 10.31876 7.987828e-17
## 10 -log(prop)/k2 0.3 46.783265 4.1532955 11.26413 9.552912e-19
## 11 -log(prop)/k3 0.3 39.685266 3.5756966 11.09861 2.064093e-18
## 12 -log(prop)/k4 0.3 55.078164 4.5913725 11.99601 3.257068e-20The differences between the half-lives (and other degradation times) can be calculated as well:
funList <- list(~ (k2 - k1)/(k1 * k2) * log(prop),
~ (k3 - k1)/(k1 * k3) * log(prop),
~ (k4 - k1)/(k1 * k4) * log(prop))
gnlht(modNlin, funList, const = data.frame(prop = c(0.7, 0.5, 0.3)))
## Form prop Estimate SE t-value p-value
## 1 (k2 - k1)/(k1 * k2) * log(prop) 0.7 5.486900 1.473859 3.722813 3.468973e-04
## 2 (k3 - k1)/(k1 * k3) * log(prop) 0.7 3.384130 1.334340 2.536183 1.297111e-02
## 3 (k4 - k1)/(k1 * k4) * log(prop) 0.7 7.944250 1.583814 5.015900 2.718444e-06
## 4 (k2 - k1)/(k1 * k2) * log(prop) 0.5 10.663013 2.864235 3.722813 3.468973e-04
## 5 (k3 - k1)/(k1 * k3) * log(prop) 0.5 6.576577 2.593100 2.536183 1.297111e-02
## 6 (k4 - k1)/(k1 * k4) * log(prop) 0.5 15.438524 3.077917 5.015900 2.718444e-06
## 7 (k2 - k1)/(k1 * k2) * log(prop) 0.3 18.521287 4.975078 3.722813 3.468973e-04
## 8 (k3 - k1)/(k1 * k3) * log(prop) 0.3 11.423287 4.504125 2.536183 1.297111e-02
## 9 (k4 - k1)/(k1 * k4) * log(prop) 0.3 26.816185 5.346236 5.015900 2.718444e-06The possibility of passing constants in a data.frame adds flexibility with respect to the ‘deltaMethod()’ function in the ‘car’ package. For example, we can use this method to make predictions:
funList <- list(~ A1 * exp (- k1 * Time), ~ A2 * exp (- k2 * Time),
~ A3 * exp (- k3 * Time), ~ A4 * exp (- k4 * Time))
pred <- gnlht(modNlin, funList, const = data.frame(Time = seq(0, 67, 1)))
head(pred)
## Form Time Estimate SE t-value p-value
## 1 A1 * exp(-k1 * Time) 0 94.83198 4.795948 19.77336 3.931106e-34
## 2 A2 * exp(-k2 * Time) 0 102.08592 4.316122 23.65223 6.671764e-40
## 3 A3 * exp(-k3 * Time) 0 99.58530 4.463418 22.31144 5.485452e-38
## 4 A4 * exp(-k4 * Time) 0 111.62446 4.183588 26.68151 5.980930e-44
## 5 A1 * exp(-k1 * Time) 1 90.87694 4.381511 20.74101 1.223613e-35
## 6 A2 * exp(-k2 * Time) 1 99.49225 4.062186 24.49229 4.617181e-41
r
tail(pred)
## Form Time Estimate SE t-value p-value
## 267 A3 * exp(-k3 * Time) 66 13.446308 2.095933 6.415427 6.838561e-09
## 268 A4 * exp(-k4 * Time) 66 26.375174 2.618594 10.072266 2.555133e-16
## 269 A1 * exp(-k1 * Time) 67 5.462339 1.363509 4.006090 1.287737e-04
## 270 A2 * exp(-k2 * Time) 67 18.202556 2.358965 7.716331 1.752360e-11
## 271 A3 * exp(-k3 * Time) 67 13.044500 2.068082 6.307534 1.106297e-08
## 272 A4 * exp(-k4 * Time) 67 25.804886 2.607131 9.897810 5.827813e-16Although this is not very fast, in contrast to the ‘predict()’ method for ‘nls’ objects, it has the advantage of reporting standard errors.
Hope this is useful. Happy coding!
Andrea Onofri
Department of Agricultural, Food and Environmental Sciences
University of Perugia (Italy)
Borgo XX Giugno 74
I-06121 - PERUGIA
References
- John Fox and Sanford Weisberg (2019). An {R} Companion to Applied Regression, Third Edition. Thousand Oaks CA:Sage. URL: https://socialsciences.mcmaster.ca/jfox/Books/Companion/
- Torsten Hothorn, Frank Bretz and Peter Westfall (2008). Simultaneous Inference in General Parametric Models. Biometrical Journal 50(3), 346–363.
- Ritz, C., Baty, F., Streibig, J. C., Gerhard, D. (2015) Dose-Response Analysis Using R PLOS ONE, 10(12), e0146021
- Vischetti, C., Marini, M., Businelli, M., Onofri, A., 1996. The effect of temperature and co-applied herbicides on the degradation rate of phenmedipham, chloridazon and metamitron in a clay loam soil in the laboratory, in: Re, A.D., Capri, E., Evans, S.P., Trevisan, M. (Eds.), “The Environmental Phate of Xenobiotics”, Proceedings X Symposium on Pesticide Chemistry, Piacenza. La Goliardica Pavese, Piacenza, pp. 287–294.